HCLSIG/LODD/Data
NOTE: WORK-IN-PROGRESS fu-berlin datasets are being hosted by Bio2RDF. Several are already there. Updates to this page and CKAN Datahub are pending..
| Name | Topic | Short Description | Size and coverage | Status / Activity | Example Instances | SPARQL Endpoint |
| DrugBank | Drugs | Drugbank.ca provides drug (i.e., chemical, pharmacological and pharmaceutical) data with comprehensive drug target (i.e., sequence, structure, and pathway) information (doi:10.1093/nar/gkj067) | 766,920 triples; 4,800 drugs, 2,500 protein sequences | updated regularly | Varenicline via Marbles, via OpenLink Data Explorer | http://www4.wiwiss.fu-berlin.de/drugbank/sparql |
| LinkedCT | Clinical Trials | Linked data source of trials from ClinicalTrials.gov | ~25 million triples, 106,000 trials (as of April 2011) | Updated automatically at all times, refer to FAQ for more details. | Breast Cancer (Condition), a NCT00999557 (Trial), Toronto (City). | http://data.linkedct.org/sparql |
| DailyMed | All FDA-approved Structured Product Labels (SPLs) for currently marketed drugs enhanced with indexing to pharmacogenomics information and NDF-RT drug class assignments | Data available via a D2R server (sample data), as an RDF dumpt (full data, ntriples), or from Virtuoso RDF Store (contact maintainer) | 1,604,893 triples, 36,000+ product labels | Updated every Thursday using information from the DailyMed RSS feed | SPL for Venlafaxine Hydrochloride (American Health Packaging) | http://purl.org/net/nlprepository/linkedSPLs |
| DBpedia | Drugs/ Diseases/ Proteins | RDF data about 2.49 million things that has been extracted from Wikipedia | 218 million RDF triples; 2,300 drugs, 2,200 proteins | updated every 3 months | Aspirin, HIV | http://dbpedia.org/sparql |
| Diseasome | Diseases / Genes | Diseasome describes characteristics of disorders and disease genes linked by known disorder–gene associations | 91,182 triples; 2,600 genes | updated 2006 | Alzheimer's via Marbles, via OpenLink Data Explorer | http://www4.wiwiss.fu-berlin.de/diseasome/sparql |
| The Drug Interaction Knowledge Base | Drugs / Metabolic Inhibition Drug-drug Interactions (DDIs) / Claims and Evidence for drug mechanisms and DDIs | A D2R server of more than 60 drugs currently in the DIKB | >41K | Updated 12/21/2012 | paroxetine, atorvastatin | http://dbmi-icode-01.dbmi.pitt.edu:2020/ |
| RDF-TCM | Genes / Diseases / Medicine / Ingredients | Traditional Chinese medicine, gene and disease association dataset and a linkset mapping TCM gene symbols to Extrez Gene IDs created by Neurocommons | 117,643 | updated August 2009 (stable) | Ginkgo biloba | http://www.open-biomed.org.uk/sparql/endpoint/tcm |
| RxNorm | Drugs | A linked version of the NLM's RxNorm database that connects prescription drugs, ingredients, and NDC through RXCUI a concept unique identifier. RxNorm is a product developed by NIH’s National Library of Medicine. It currently interlinks 12 different drug vocabularies around a unique concept identifier. Due to licensing only six of the drug vocabularies are made available as part of the LODD cloud. This includes: Medical Subject Headings,, Metathesaurus FDA National Drug Code Directory, Metathesaurus FDA Structured Product Labels, National Drug File, RxNorm Vocabulary, Veterans Health Administration National Drug File
Links are provided connecting RxNorm to drug bank and to the UMLS. |
over 7.7 million triples; 165,806 RXCUI (Concept Unique Identifiers) Unique drugs and ingredients; 332,754 RXAUI (Atomic Unique Identifiers) sourced terms | Based on 3/2010 Rxnorm Release; Last updated 5/2010 | Singulair from the Metathesaurus FDA Structured Product Labels | http://link.informatics.stonybrook.edu/sparql/ |
| SIDER | Diseases / Side Effects | SIDER contains information on marketed drugs and their adverse effects (doi:10.1038/msb.2009.98) | 192,515 triples; 63,000 adverse effect reports, 1,737 genes | updated 2009 | Confusion via Marbles | http://www4.wiwiss.fu-berlin.de/sider/sparql |
| STITCH | Chemicals / Proteins | STITCH contains information on chemicals, proteins, and their interactions (doi:10.1093/nar/gkm795) | 7,500,000 chemicals; 500,000 proteins; 370 organisms | updated July 2009 | Lactose via Marbles | http://www4.wiwiss.fu-berlin.de/stitch/sparql |
| Medicare | Medicare Formulary | xxx | xxx | xxx | xxx | http://www4.wiwiss.fu-berlin.de/medicare/sparql |
| ChEMBL | Chemical / Assays (Proteins, Organisms) / Papers | ChEMBL contains information on trial drugs with information about activity against targets like but not limited to proteins. All is backed up by and linked to literature. Includes links to Bio2RDF for ChEBI and Uniprot. License: CC-BY-SA. | ~130M triples | Updated 2010-01 | A IC50 activity. | http://rdf.farmbio.uu.se/chembl/sparql |
| WHO's Global Health Observatory (GHO) | Infectious Diseases /Demography / Socioeconomic Conditions / Environmental Factors | Data and statistics for infectious diseases at country, regional, and global levels | ~3M triples | Updated 2012-05 | xxx | http://gho.aksw.org |
| University of Pittsburgh NLP Repository | Drugs / Procedures / Diagnoses | A semantic index of concepts present in 800 full-text clinical notes from the University of Pittsburgh NLP Repository | 38.664 | Proof of concept -- Updated 02/25/2011 | Concepts from a sample radiology report | http://dbmi-icode-01.dbmi.pitt.edu:8080/sparql |
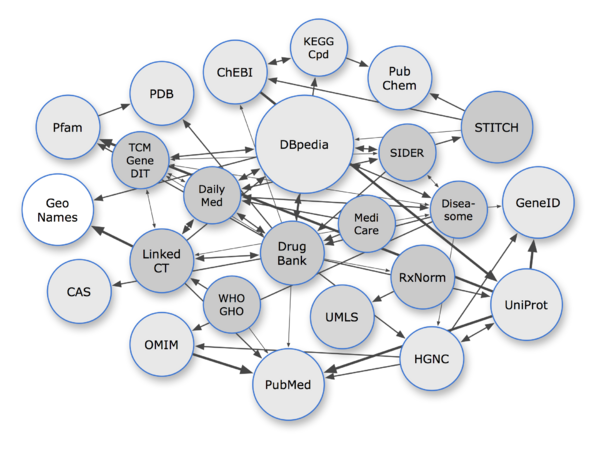
A graph of some of the LODD datasets (dark grey), related biomedical datasets (light grey), related general-purpose datasets (white) and their interconnections. Line weights correspond to the number of links. The direction of an arrow indicates the dataset that contains the links, e.g., an arrow from A to B means that dataset A contains RDF triples that use identifiers from B. Bidirectional arrows usually indicate that the links are mirrored in both datasets. More on the interlinking methodology and statistics can be found on the Interlinking page.
The LODD datasets have been crawled by the SWSE Semantic Web search engine and can be accessed via a faceted browsing interface at [1] (Example query: Varenicline).
Most of the LODD datasets have also been integrated into the SPARQL endpoint of the HCLS Knowledge Base, see the wiki page of the HCLS KB for further information.
Bio2RDF Data Sets
The Bio2RDF project has published 40 biology-, gene- and medical-related datasets (altogether 2.3 billion triples). The datasets are available via SPARQL endpoints and as Linked Data. It is recommended that you use the Bio2RDF Java Servlet, and optionally download the databases for efficient personal use. Running your own instance of the OpenLink Virtuoso AMI for EC2 is also an option (and for basic URI resolution doesn't require the Java Servlet, although if you want advanced queries you should still download it and configure it to query your EC2 sparql endpoint).
- Bio2RDF sparql endpoint list Sparql endpoint list in RDF
- Identification of an autoimmune enteropathy-related 75-kilodalton antigen, via an OpenLink hosted edition of Bio2Rdf
- Structure of the gene encoding the human cyclin-dependent kinase inhibitor p18 and mutational analysis in breast cancer, via an OpenLink hosted edition of Bio2Rdf
- PubMed article viewed using the Marbles Linked Data browser.
- PubMed author viewed using the Marbles Linked Data browser.
- OMIM Killer Cell Lectin-Like Receptor viewed using the Marbles Linked Data browser.
- Falcons Search for KILLER CELL. The Bio2RDF data has been crawled by the Falcons Semantic Web Search engine. This is an example on how the data is accessed by humans using the search engine. Falcons also offers an API that can by used by applications to access the data.
Chem2bio2RDF
- Information about the chem2bio2rdf data sets
Data Sets for the LODD Task
To complement the drug-related Web of Data build by the LODD effort, the following data sets could/should also be published as Linked Data.
The LODD effort is currently gathering more information about relevant datasets. See also Evaluation of LODD Data Sets for current evaluation results.
- Adis R&D Insight
- chEBI
- ChemBlast
- ChemSpider
- ClinicalTrials.gov
- Citeline TrialTrove
- DailyMed
- DBpedia
- Diseasome
- Drug Bank
- DrugDB
- Drugome
- Drug Ontology
- Investigational Drug Database - Proprietary
- IMS
- KEGG Drug
- LillyTrials
- MedWatch
- National Drug Code
- OMIM
- Orange Book
- Pharmaprojects - Proprietary
- PubChem
- RxNorm
- VA NDF-RT
- Other data sources could include blogs, discussion boards, wikis, etc.
- and....
Alternative Herbal Medicine use case
Identified Based Linkage Points
Data Set Attributes
- Licensing
- Data Format
- Identifiers