












Data utility depends on whether it
Data dimensions
Size/Structure (false) dichotomy
Some day, these will be views on the same data... we hope.


practice data +
to build

inclusion/exclusion criteria

protocol specifies a small number of specific acquisitions with special terminology codes.
Too early to tell.
263870008 (SNOMED code for Sequestration)
Around 2009, CDISC members identified a need for intra and inter pharma sharing of trial metadata:
They were going to call it SHARE.
4 years later,
Reduce the n2 pressure.
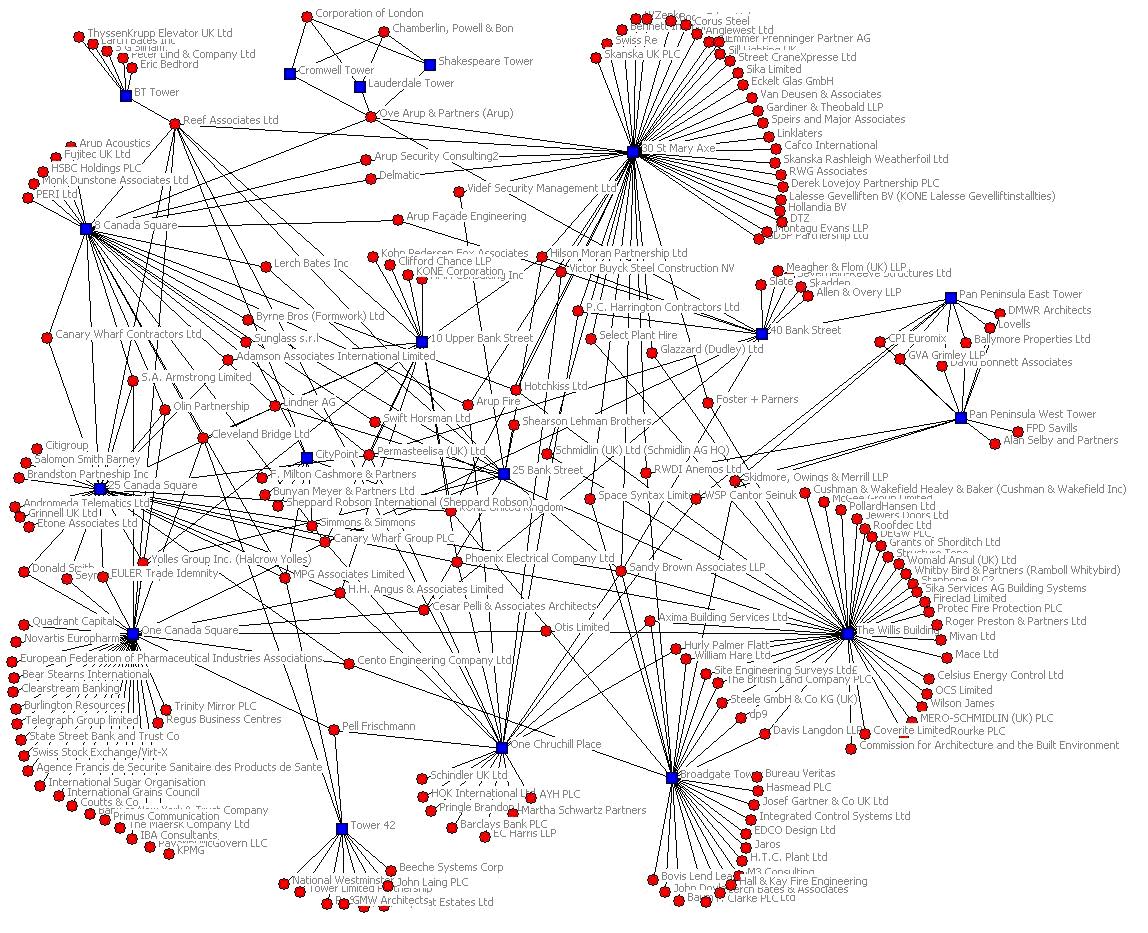
Example detailed clinical observation:
"Information" model sounds presumptuous;
it's just a term of art.
Separate network of relationship from the values
Info model still holds relationships semantics.
a clinical observation
a procedure
a measurement
condition severity
prescription fulfillment
assertion that some measurement is a
reference range for another.
person
organization
device
medication.
For example, Dr. Leonid Rogozov participated
in an emergency appendectomy as a surgeon,
patient, and, if we was diligent,
records custodian.
Structural attributes, e.g. mood:
considered, planned, accomplished
Factors commonalities in e.g. Prescription vs. Injection.
Example RMIM using the conventional colors

Our earlier example is just Acts and ActRelationships. The rimITS provides a straightforward XML representation of RIM:
RIM ITS XML
<content xsi:type="Observation"
classCode="OBS" moodCode="EVN">
...
<value code="241938005"
displayName="penicillin-induced
anaphylaxis (disorder)" .../>
<inboundRelationship typeCode="SUBJ">
<source ...>
<code code="SEV" displayName="Severity Observation" .../>
<value code="423132009"
displayName="Grade 4/5"/>
</source> </inboundRelationship>
<outboundRelationship typeCode="EVID">...
<target ...>
<code code="ASSERTION" .../>
<value code="247472004"
displayName="weal (disorder)"/>
</target> </outboundRelationship>
... <!-- etc. for 373895009 (acute respiratory distress) and 45007003 (low blood pressure) -->
</content>
<rim-graph xmlns="urn:hl7-org:v3-rim" xmlns:xsi="http://www.w3.org/2001/XMLSchema-instance" version="0229">
<content xsi:type="Observation" classCode="OBS" moodCode="EVN">
<code code="ASSERTION" displayName="Assertion"
codeSystem="2.16.840.1.113883.5.4" codeSystemName="HL7 ActCode"/>
<value xsi:type="CD" code="241938005" displayName="penicillin-induced anaphylaxis (disorder)"
codeSystem="2.16.840.1.113883.6.96" codeSystemName="SNOMED CT"/>
<inboundRelationship typeCode="SUBJ">
<source xsi:type="Observation" classCode="OBS" moodCode="EVN">
<code code="SEV" displayName="Severity Observation"
codeSystem="2.16.840.1.113883.5.4" codeSystemName="HL7 ActCode"/>
<value xsi:type="CD" code="423132009"
codeSystem="2.16.840.1.113883.6.96" codeSystemName="SNOMED CT"/>
</source>
</inboundRelationship>
<outboundRelationship typeCode="EVID">
<target xsi:type="Observation" classCode="OBS" moodCode="EVN">
<code code="ASSERTION" displayName="Assertion"
codeSystem="2.16.840.1.113883.5.4" codeSystemName="HL7 ActCode"/>
<value xsi:type="CD" code="247472004" displayName="weal (disorder)"
codeSystem="2.16.840.1.113883.6.96" codeSystemName="SNOMED CT"/>
</target>
</outboundRelationship>
<outboundRelationship typeCode="EVID">
<target xsi:type="Observation" classCode="OBS" moodCode="EVN">
<code code="ASSERTION" displayName="Assertion"
codeSystem="2.16.840.1.113883.5.4" codeSystemName="HL7 ActCode"/>
<value xsi:type="CD" code="373895009" displayName="acute respiratory distress"
codeSystem="2.16.840.1.113883.6.96" codeSystemName="SNOMED CT"/>
</target>
</outboundRelationship>
<outboundRelationship typeCode="EVID">
<target xsi:type="Observation" classCode="OBS" moodCode="EVN">
<code code="ASSERTION" displayName="Assertion"
codeSystem="2.16.840.1.113883.5.4" codeSystemName="HL7 ActCode"/>
<value xsi:type="CD" code="45007003" displayName="low blood pressure (disorder)"
codeSystem="2.16.840.1.113883.6.96" codeSystemName="SNOMED CT"/>
</target>
</outboundRelationship>
</content>
</rim-graph>
General-purpose graph structure for medical information:
Tells you where to write but not how to write:
information model (RIM)
Taxonomies used for:
terminology model
<entry typeCode="DRIV">
<act classCode="ACT" moodCode="EVN">
<templateId root="2.16.840.1.113883.10.20.22.4.30"/>
<id root="69724b9d-5e77-402f-a5b6-3379dde3e8a9"/>
<code code="48765-2" displayName="ALLERGIES, ADVERSE REACTIONS, ALERTS" codeSystem="2.16.840.1.113883.6.1" codeSystemName="LOINC"/>
<statusCode code="active"/>
<effectiveTime>
<low value="20120806"/>
</effectiveTime>
<entryRelationship typeCode="SUBJ">
<observation classCode="OBS" moodCode="EVN">
<templateId root="2.16.840.1.113883.10.20.22.4.7"/>
<id root="69724b9d-5e77-402f-a5b6-3379dde3e8a9"/>
<code code="ASSERTION" codeSystem="2.16.840.1.113883.5.4" codeSystemName="HL7ActCode"/>
<statusCode code="completed"/>
<effectiveTime>
<low value="20120806"/>
<high nullFlavor="UNK"/>
</effectiveTime>
<value code="416098002" displayName="drug allergy" codeSystem="2.16.840.1.113883.6.96" codeSystemName="SNOMED CT" xsi:type="CD"/>
<participant typeCode="CSM">
<participantRole classCode="MANU">
<playingEntity classCode="MMAT">
<code code="2670" displayName="Codeine" codeSystem="2.16.840.1.113883.6.88" codeSystemName="RxNorm">
<originalText>
<reference value="#AllergyDescription_0"/>
</originalText>
<translation code="1550" displayName="Codeine" codeSystem="2.16.840.1.113883.4.65" codeSystemName="FDB Hierarchical Ingredient Code Sequence Number (HIC_SEQN)"/>
</code>
<name>Codeine </name>
</playingEntity>
</participantRole>
</participant>
<entryRelationship typeCode="MFST" inversionInd="true">
<observation classCode="OBS" moodCode="EVN">
<templateId root="2.16.840.1.113883.10.20.22.4.9"/>
<id root="69724b9d-5e77-402f-a5b6-3379dde3e8a9"/>
<code code="ASSERTION" codeSystem="2.16.840.1.113883.5.4" codeSystemName="HL7ActCode"/>
<text>
<reference value="#AllergyReaction_0"/>
</text>
<statusCode code="completed"/>
<effectiveTime>
<low value="20120806"/>
</effectiveTime>
<value xsi:type="CD" nullFlavor="UNK">
<originalText>
<reference value="#AllergyReaction_0"/>
</originalText>
</value>
</observation>
</entryRelationship>
<entryRelationship typeCode="SUBJ" inversionInd="true">
<observation classCode="OBS" moodCode="EVN">
<templateId root="2.16.840.1.113883.10.20.22.4.28"/>
<code code="33999-4" displayName="STATUS" codeSystem="2.16.840.1.113883.6.1" codeSystemName="LOINC"/>
<text>
<reference value="#AllergyStatus_0"/>
</text>
<statusCode code="completed"/>
<value code="55561003" displayName="active" codeSystem="2.16.840.1.113883.6.96" codeSystemName="SNOMED CT" xsi:type="CE"/>
</observation>
</entryRelationship>
<entryRelationship typeCode="REFR">
<act classCode="ACT" moodCode="EVN">
<templateId root="1.3.6.1.4.1.19376.1.5.3.1.4.4.1"/>
<id extension="a778b36c-2a08-4f41-bc2a-3f29293c4e3c" root="2.201"/>
<code nullFlavor="UNK"/>
</act>
</entryRelationship>
</observation>
</entryRelationship>
</act>
</entry>
[ a rim:ActRelationship ; rim:ActRelationship.typeCode [ hl7:coding [ dt:CDCoding.code "DRIV" ; dt:CDCoding.codeSystem "???" ] ] ; rim:ActRelationship.source _:section0 ; rim:ActRelationship.target _:section0_allergyProblemAct0 ] . _:section0_allergyProblemAct0 a rim:Observation ; rim:Act.id [ a dt:DSET_II ; dt:COLL.item [ dt:II.root "69724b9d-5e77-402f-a5b6-3379dde3e8a9" ] ] ; rim:Act.classCode [ hl7:coding [ dt:CDCoding.code "ACT" ; dt:CDCoding.codeSystem "???" ] ] ; rim:Act.moodCode [ hl7:coding [ dt:CDCoding.code "EVN" ; dt:CDCoding.codeSystem "2.16.840.1.113883.5.1001" ] ] ; rim:Act.effectiveTime-low "2012-08-06"^^xsd:date ; rim:Act.statusCode "active" ; rim:Act.code [ hl7:coding [ dt:CDCoding.code "48765-2" ; dt:CDCoding.codeSystem "2.16.840.1.113883.6.1" ; dt:CDCoding.displayName "ALLERGIES, ADVERSE REACTIONS, ALERTS" ; dt:CDCoding.codeSystemName "LOINC" ] ]; rim:Observation.value [ ] .# SUBJ [ a rim:ActRelationship ; rim:ActRelationship.typeCode [ hl7:coding [ dt:CDCoding.code "REFR" ; dt:CDCoding.codeSystem "???" ] ] ; rim:ActRelationship.source _:section0_allergyProblemAct0 ; rim:ActRelationship.target _:section0_allergyProblemAct0_entryRelationship0 ] . _:section0_allergyProblemAct0_entryRelationship0 a rim:Observation ; rim:Act.id [ a dt:DSET_II ; dt:COLL.item [ dt:II.root "69724b9d-5e77-402f-a5b6-3379dde3e8a9" ] ] ; rim:Act.classCode [ hl7:coding [ dt:CDCoding.code "OBS" ; dt:CDCoding.codeSystem "2.16.840.1.113883.5.6" ] ] ; rim:Act.moodCode [ hl7:coding [ dt:CDCoding.code "EVN" ; dt:CDCoding.codeSystem "2.16.840.1.113883.5.1001" ] ] ; rim:Act.effectiveTime-low "2012-08-06"^^xsd:date ; rim:Act.statusCode "completed" ; rim:Act.code [ hl7:coding [ dt:CDCoding.code "ASSERTION" ; dt:CDCoding.codeSystem "2.16.840.1.113883.5.4" ; dt:CDCoding.codeSystemName "HL7ActCode" ] ]; rim:Observation.value [ a dt:PQ ; dt:CD.code [ a dt:CD ; hl7:coding [ rim:CDCoding.code "416098002" ; rim:CDCoding.codeSystem "2.16.840.1.113883.6.96" ; rim:CDCoding.displayName "drug allergy" ; rim:CDCoding.codeSystemName "SNOMED CT" ] ] ] .# Participant [ a rim:Participation ; rim:Participation.typeCode [ hl7:coding [ dt:CDCoding.code "CSM" ; dt:CDCoding.codeSystem "???" ] ] ; rim:Participation.act _:section0_allergyProblemAct0_entryRelationship0 ; rim:Participation.role _:section0_allergyProblemAct0_entryRelationship0_participant0 ] . _:section0_allergyProblemAct0_entryRelationship0_participant0 a rim:Role ; rim:Role.player _:section0_allergyProblemAct0_entryRelationship0_participant0_player ; rim:Role.scoper _:section0_allergyProblemAct0_entryRelationship0_participant0_scoper ; rim:Role.classCode [ hl7:coding [ dt:CDCoding.code "ROL" ; dt:CDCoding.codeSystem "???" ] ] ; . _:section0_allergyProblemAct0_entryRelationship0_participant0_player a rim:Person ; rim:Entity.classCode [ hl7:coding [ dt:CDCoding.code "PSN" ; dt:CDCoding.codeSystem "???" ] ] ; rim:Entity.determinerCode [ hl7:coding [ dt:CDCoding.code "INSTANCE" ; dt:CDCoding.codeSystem "???" ] ] ; rim:Entity.name [ a dt:COLL_EN ; ]; ; ;
RDF representation uses bridg:Class.relationship.Class notation
to capture relationships between classes (UML boxes).
Navigated as UML diagrams and "instance" diagrams
or...
Portable BRIDG Representation
Terminology Serves a variety of purposes:
The presence of a terminology (aka. "Ontology" when used in the broader sense) means little or nothing. You've got some codes (URI's) that you can embed in data or link to, but unless you know the scope and purpose of the terminology, it is useless at best, and can actually be quite dangerous.
As an example, scroll to the bottom of: the MeSH definition of Neoplasms. Note that (implicitly) Neoplastic process, Precancerous conditions and Neoplastic pregnancy complications are all "narrower" than Neoplasms. This (obviously) does not mean, however, that an instance of a Precancerous condition is an instance of a Neoplasm
Going in the other direction, look at the definition of Appendicitis in SNOMED CT. The (implicit) assertion is that:
sctfsn:Appendicitis rdfs:subClassOf sctfsn:Disorder_of_appendix.
states that any instance of Appendicitis is also necessarily an instance of a Disorder of Appendix.
The asssertion:sctfsn:Appendicitis skos:broader sctfsn:Disorder_of_appendix.
only makes a very fuzzy assertion that "conceptually" whenever you think of Appendicitis, you should probably also be thinking of Disorder of Appendix.
Similarly, if you try to use a classification system such as ICD-10 to infer knowledge about the domain of medicine, you will find yourself reaching some rather startling conclusions, such as the "fact" that Neonatal diabetes mellitus is not a type of Diabetes Mellitus. The correct interpretation can be nothing more than:
The World Health Organization has determined that instances of Neonatal diabetes will be categorized as Transitory disorders of carbohydrate metabolism specific to fetus and newborn
Resources/StatedRelationshipsToOwlKRSS directoryperl tls2_StatedRelationshipsToOwlKRSS_INT_20150731.pl OWLF ../../Snapshot/.
(Note that this does Stated relationships. Other options are possible...)or use a specialized form
or
git@github.com:w3c/hcls-fhir-rdf.gitwget https://hl7-fhir.github.io/observation-example.xmlxsltproc -stringparam fhirdefs ../data/definitions.xml xsl/transform.xsl observation-example.xmlsimilar to FHIR, but using the CDA-Turtle XSLT (see online example)
_:section0_allergyProblemAct0 a rim:Observation ; rim:Act.id [ a dt:DSET_II ; dt:COLL.item [ dt:II.root "69724b9d-5e77-402f-a5b6-3379dde3e8a9" ] ] ; rim:Act.classCode [ hl7:coding [ dt:CDCoding.code "ACT" ; dt:CDCoding.codeSystem "???" ] ] ; rim:Act.moodCode [ hl7:coding [ dt:CDCoding.code "EVN" ; dt:CDCoding.codeSystem "2.16.840.1.113883.5.1001" ] ] ; rim:Act.effectiveTime-low "2012-08-06"^^xsd:date ; rim:Act.statusCode "active" ; rim:Act.code [ hl7:coding [ dt:CDCoding.code "48765-2" ; dt:CDCoding.codeSystem "2.16.840.1.113883.6.1" ; dt:CDCoding.displayName "ALLERGIES, ADVERSE REACTIONS, ALERTS" ; dt:CDCoding.codeSystemName "LOINC" ] ]; rim:Observation.value [ ] .# SUBJ [ a rim:ActRelationship ; rim:ActRelationship.typeCode [ hl7:coding [ dt:CDCoding.code "REFR" ; dt:CDCoding.codeSystem "???" ] ] ; rim:ActRelationship.source _:section0_allergyProblemAct0 ; rim:ActRelationship.target _:section0_allergyProblemAct0_entryRelationship0 ] . _:section0_allergyProblemAct0_entryRelationship0 a rim:Observation ; rim:Act.id [ a dt:DSET_II ; dt:COLL.item [ dt:II.root "69724b9d-5e77-402f-a5b6-3379dde3e8a9" ] ] ; rim:Act.classCode [ hl7:coding [ dt:CDCoding.code "OBS" ; dt:CDCoding.codeSystem "2.16.840.1.113883.5.6" ] ] ; rim:Act.moodCode [ hl7:coding [ dt:CDCoding.code "EVN" ; dt:CDCoding.codeSystem "2.16.840.1.113883.5.1001" ] ] ; rim:Act.effectiveTime-low "2012-08-06"^^xsd:date ; rim:Act.statusCode "completed" ; rim:Act.code [ hl7:coding [ dt:CDCoding.code "ASSERTION" ; dt:CDCoding.codeSystem "2.16.840.1.113883.5.4" ; dt:CDCoding.codeSystemName "HL7ActCode" ] ]; rim:Observation.value [ a dt:PQ ; dt:CD.code [ a dt:CD ; hl7:coding [ rim:CDCoding.code "416098002" ; rim:CDCoding.codeSystem "2.16.840.1.113883.6.96" ; rim:CDCoding.displayName "drug allergy" ; rim:CDCoding.codeSystemName "SNOMED CT" ] ] ] .# Participant [ a rim:Participation ; rim:Participation.typeCode [ hl7:coding [ dt:CDCoding.code "CSM" ; dt:CDCoding.codeSystem "???" ] ] ; rim:Participation.act _:section0_allergyProblemAct0_entryRelationship0 ; rim:Participation.role _:section0_allergyProblemAct0_entryRelationship0_participant0 ] . _:section0_allergyProblemAct0_entryRelationship0_participant0 a rim:Role ; rim:Role.player _:section0_allergyProblemAct0_entryRelationship0_participant0_player ; rim:Role.scoper _:section0_allergyProblemAct0_entryRelationship0_participant0_scoper ; rim:Role.classCode [ hl7:coding [ dt:CDCoding.code "ROL" ; dt:CDCoding.codeSystem "???" ] ] ; . _:section0_allergyProblemAct0_entryRelationship0_participant0_player a rim:Person ; rim:Entity.classCode [ hl7:coding [ dt:CDCoding.code "PSN" ; dt:CDCoding.codeSystem "???" ] ] ; rim:Entity.determinerCode [ hl7:coding [ dt:CDCoding.code "INSTANCE" ; dt:CDCoding.codeSystem "???" ] ] ; rim:Entity.name [ a dt:COLL_EN ; ].

<ObservationShape> {
a (fhir:Observation)?,
fhir:Observation.id @<LitShape>?,
fhir:Observation.meta @<MetaShape>?,
fhir:Observation.implicitRules @<LitShape>?,
fhir:Observation.language @<LitShape>?,
fhir:Observation.text @<NarrativeShape>?,
fhir:Observation.contained @<ResourceShape>*,
fhir:Observation.extension @<ExtensionShape>*,
fhir:Observation.modifierExtension @<ExtensionShape>*,
fhir:Observation.code @<CodeableConceptShape>,
fhir:Observation.valueQuantity @<QuantityShape>?,
fhir:Observation.valueCodeableConcept @<CodeableConceptShape>?,
fhir:Observation.valuestring @<LitShape>?,
fhir:Observation.valueRange @<RangeShape>?,
fhir:Observation.valueRatio @<RatioShape>?,
fhir:Observation.valueSampledData @<SampledDataShape>?,
fhir:Observation.valueAttachment @<AttachmentShape>?,
fhir:Observation.valuetime @<LitShape>?,
fhir:Observation.valuedateTime @<LitShape>?,
fhir:Observation.valuePeriod @<PeriodShape>?,
fhir:Observation.dataAbsentReason @<CodeableConceptShape>?,
fhir:Observation.interpretation @<CodeableConceptShape>?,
fhir:Observation.comments @<LitShape>?,
fhir:Observation.appliesdateTime @<LitShape>?,
fhir:Observation.appliesPeriod @<PeriodShape>?,
fhir:Observation.issued @<LitShape>?,
fhir:Observation.status @<LitShape>,
fhir:Observation.reliability @<LitShape>?,
fhir:Observation.bodySiteCodeableConcept @<CodeableConceptShape>?,
fhir:Observation.bodySiteReference @<ReferenceShape>?,
fhir:Observation.method @<CodeableConceptShape>?,
fhir:Observation.identifier @<IdentifierShape>*,
fhir:Observation.subject @<ReferenceShape>?,
fhir:Observation.specimen @<ReferenceShape>?,
fhir:Observation.performer @<ReferenceShape>*,
fhir:Observation.device @<ReferenceShape>?,
fhir:Observation.encounter @<ReferenceShape>?,
fhir:Observation.referenceRange @<BackboneElementShape>*,
fhir:Observation.related @<BackboneElementShape>*
}
FHIR has a REST-based HTTP protocol
hence the Search Parameters
much more detailed that anything in LDP.
FDA (CDER) project billed as:
Establish common study data standards:
Code everything!
minimize:
cope gracefully with:
interoperate with EHRS:
Existing Models/Standards
Controlled Terminologies:
bridg:PerformedObservation

Portable BRIDG Representation
:SerumCreatinineLevel a owl:Class ; rdfs:subClassOf [ owl:onProperty bridg:resultIn ; owl:allValuesFrom bridg:PerformedClinicalResult ], [ owl:onProperty bridg:resultIn ; owl:cardinality 1 ] .
:GraftBPARAssessment a owl:Class ; rdfs:subClassOf core:NegativeOutcome ; owl:equivalentClass [ a owl:Class ; owl:intersectionOf ( [ a owl:Restriction ; owl:onProperty core:afterIntervention ; owl:someValuesFrom [ a owl:Restriction ; owl:onProperty core:hasPathologyFinding ; owl:hasValue :BanffIII ] ] [ a owl:Restriction ; owl:onProperty core:hasResultValue ; owl:hasValue :NonFunctioningGraft ] ) ] .
<http://www.w3.org/2013/12/FDA-TA/RenalTransplantation> a owl:Ontology ; owl:imports <http://www.w3.org/2013/12/FDA-TA/core> , <http://www.w3.org/2013/12/FDA-TA/renal> , <http://www.w3.org/2013/12/FDA-TA/transplant> .
SELECT ?outcomeType ?dose
(AVG(?endpoint-time) AS ?rate)
WHERE {
# drug of interest
?adminDrug dt:CD.displayName "Upsidasium" ;
… codingSystem … .
# subjects in studies about drug
?arm :studySubject ?adminDrug ;
:studyParticipation ?subject .
# demographic selection
?subject :taxon ncbitax:7609 .
# outcomes assessing prescription performance
?outcome :intervention ?p ;
:value [ a ?outcomeType ] .
# ... of that drug on that subject (participation)
?p a :Prescription ;
:involvedSubject ?subject ;
:medication ?adminDrug .
} ORDER BY ?dose
:subjectspostOpDay3GFR a rrej:RenalFunctionObservation ;
mm:observationTime "2013-07-08T14:50:00"^^xsd:dateTime ;
rrej:gfrFlowRate [ data:value 12.0 ; data:units ucum:mL-per-minute ].
Would be in a table with codes in an EMR:
| obs id | when | coding system | ObsCode | value | units | addressing…performer… |
|---|---|---|---|---|---|---|
| 1234 | 2013-07-08T14:50:00 | CPT | 82565 | 12 | mL-per-minute | |
| 5678 | 2013-07-08T14:50:00 | SNOMED 2014-01-01 | 241374009 | 12 | mL-per-minute | |
| 4321 | 2013-07-08T14:50:00 | LOINC | 48643-1 | 12 | mL-per-minute |
or C-CDA:
<observation classCode="OBS" moodCode="EVN"> <code code="48643-1" codeSystem="2.16.840.1.113883.6.1" codeSystemName="LOINC" displayName="Glomerular filtration rate/1.73 sq M.predicted.black" /> <effectiveTime value="201307081450"/> <value type="PQ" value="12.0" unit="mL/min"/> <interpretationCode/> <referenceRange /> </observation>
Information model efforts
Needed: EPIC Cerner SIEMENS AllScripts HP athenahealth NextGen eClinicalWorks McKesson Greenway Praxis e_MDCureMD MediTab NextTech ADP ...


The popular CD datatype has:
PREFIX snomed: <http://ihtsdo.org…20130131…> PREFIX hl7: <http://hl7.org/owl/metadata#> <myObs> hl7:coding snomed:282291009 .

These are all sibling children of Degree findings:
Separate augmenters and decreasers without requiring SNOMED to do the same.
terminology model
augmented terminology

Technical barriers: