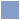













Data utility depends on whether it
Data dimensions
Size/Structure (false) dichotomy
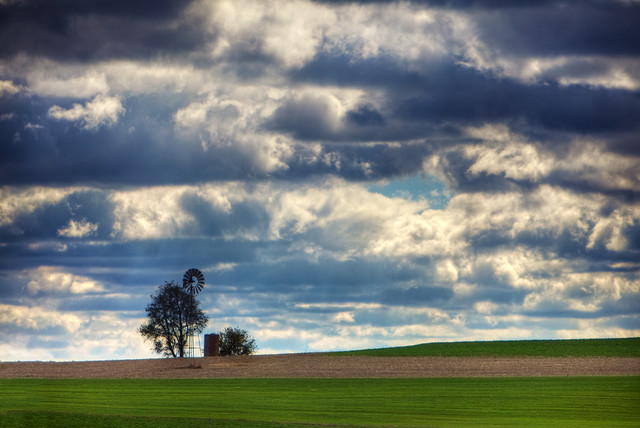


practice data +
to build

inclusion/exclusion criteria

protocol specifies a small number of specific acquisitions with special terminology codes.
Too early to tell.
263870008 (SNOMED code for Sequestration)
Around 2009, CDISC members identified a need for intra and inter pharma sharing of trial metadata:
They were going to call it SHARE.
4 years later,
General-purpose graph structure for medical information:
Tells you where to write but not how to write:
information model (RIM)
Taxonomies used for:
terminology model
Separation:
Reduce the n2 pressure.
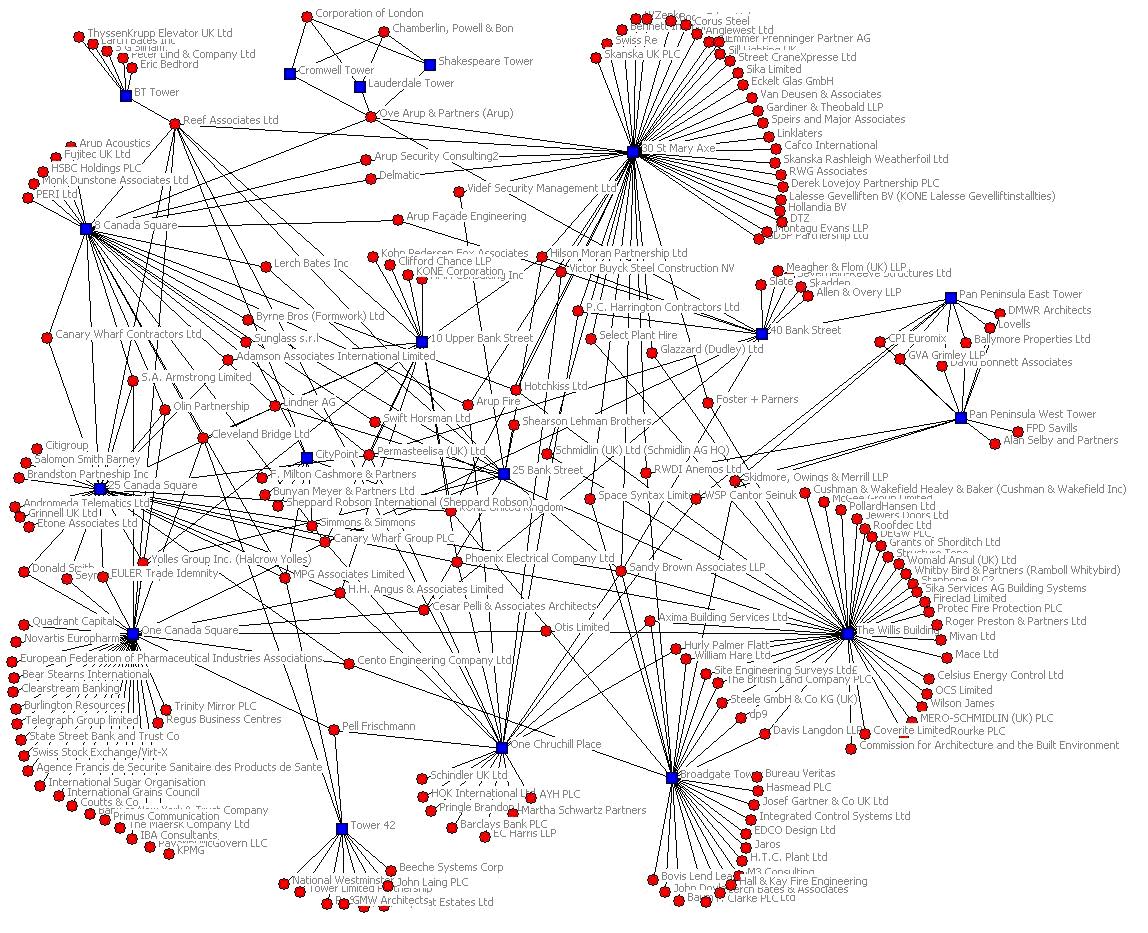
Technical barriers:
| Learn | Invent | |
|---|---|---|
| idiom | embrace | extend |
| costs | apparent | hidden |
SemWeb makes invention too easy and safe.
Try stuff out, ask questions, fix other peoples' stuff.
When I understand a model well enough to criticize it,
only then do I understand it well enough to use it.


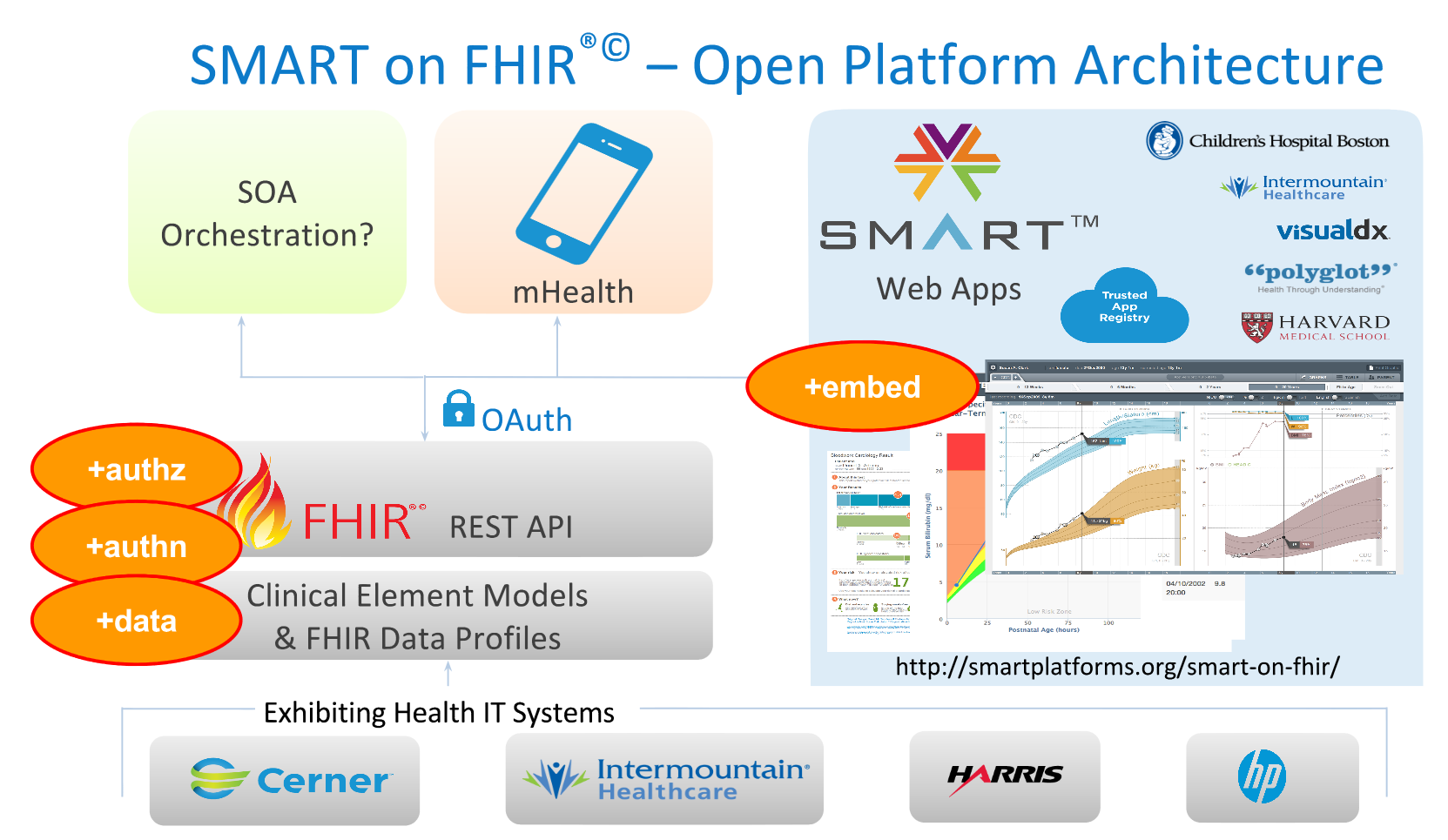
FDA (CDER) project billed as:
Establish common study data standards:
Code everything!
minimize:
cope gracefully with:
interoperate with EHRS:
Existing Models/Standards
Controlled Terminologies:
bridg:PerformedObservation
Portable BRIDG Representation
:SerumCreatinineLevel a owl:Class ; rdfs:subClassOf [ owl:onProperty bridg:resultIn ; owl:allValuesFrom bridg:PerformedClinicalResult ], [ owl:onProperty bridg:resultIn ; owl:cardinality 1 ] .
:GraftBPARAssessment a owl:Class ; rdfs:subClassOf core:NegativeOutcome ; owl:equivalentClass [ a owl:Class ; owl:intersectionOf ( [ a owl:Restriction ; owl:onProperty core:afterIntervention ; owl:someValuesFrom [ a owl:Restriction ; owl:onProperty core:hasPathologyFinding ; owl:hasValue :BanffIII ] ] [ a owl:Restriction ; owl:onProperty core:hasResultValue ; owl:hasValue :NonFunctioningGraft ] ) ] .
<http://www.w3.org/2013/12/FDA-TA/RenalTransplantation> a owl:Ontology ; owl:imports <http://www.w3.org/2013/12/FDA-TA/core> , <http://www.w3.org/2013/12/FDA-TA/renal> , <http://www.w3.org/2013/12/FDA-TA/transplant> .
SELECT ?outcomeType ?dose
(AVG(?endpoint-time) AS ?rate)
WHERE {
# drug of interest
?adminDrug dt:CD.displayName "Upsidasium" ;
… codingSystem … .
# subjects in studies about drug
?arm :studySubject ?adminDrug ;
:studyParticipation ?subject .
# demographic selection
?subject :taxon ncbitax:7609 .
# outcomes assessing prescription performance
?outcome :intervention ?p ;
:value [ a ?outcomeType ] .
# ... of that drug on that subject (participation)
?p a :Prescription ;
:involvedSubject ?subject ;
:medication ?adminDrug .
} ORDER BY ?dose
:subjectspostOpDay3GFR a rrej:RenalFunctionObservation ;
mm:observationTime "2013-07-08T14:50:00"^^xsd:dateTime ;
rrej:gfrFlowRate [ data:value 12.0 ; data:units ucum:mL-per-minute ].
Would be in a table with codes in an EMR:
| obs id | when | coding system | ObsCode | value | units | addressing…performer… |
|---|---|---|---|---|---|---|
| 1234 | 2013-07-08T14:50:00 | CPT | 82565 | 12 | mL-per-minute | |
| 5678 | 2013-07-08T14:50:00 | SNOMED 2014-01-01 | 241374009 | 12 | mL-per-minute | |
| 4321 | 2013-07-08T14:50:00 | LOINC | 48643-1 | 12 | mL-per-minute |
or C-CDA:
<observation classCode="OBS" moodCode="EVN"> <code code="48643-1" codeSystem="2.16.840.1.113883.6.1" codeSystemName="LOINC" displayName="Glomerular filtration rate/1.73 sq M.predicted.black" /> <effectiveTime value="201307081450"/> <value type="PQ" value="12.0" unit="mL/min"/> <interpretationCode/> <referenceRange /> </observation>
Information model efforts
Needed: EPIC Cerner SIEMENS AllScripts HP athenahealth NextGen eClinicalWorks McKesson Greenway Praxis e_MDCureMD MediTab NextTech ADP ...



 ?
? !
!