March 2011

Commercial semantic technologies are solving critical Healthcare and Life Sciences (HCLS) challenges. Semantic software and related integration and analysis methods are widely applied in production settings for data access, translation, semantic integration, analysis and screening, to deliver personalized medicine outcomes in research and clinical settings. The foundation for interoperable semantic technology is based on standards developed by the World Wide Web Consortium (W3C), such as the Resource Description Framework (RDF) and the Web Ontology Language (OWL). While traditional, relational data warehousing and federation approaches can scale well and are effective for many core data storage and access requirements, such approaches often fail when facing the dynamic changes and the inherent complexity of data integration requirements for Healthcare / Life Sciences (HCLS) research. Semantic integration methods assure coherence, harmonize synonyms and different terminologies, and provide an extensible, flexible data integration platform and interactive knowledge base for relevant network analysis. This paper describes the success of using an innovative semantic approach towards integration of all experimental, internal, external, clinical and public data sources. The resulting visual exploration of the integrated graph environment and the construction of characteristic marker patterns or molecular signatures are applicable to predictive functional biology-based decision support for complex translational research and personalized medicine applications. SPARQL queries can be captured visually and saved in arrays representative of specific biological functions. Being able to create, visualize and test complex models in an easy, automated way makes these methods widely applicable.
In this use case, IO Informatics’ Sentient™ semantic software technology applies patterns (extended query arrays) of combined molecular biomarkers as predictive network models. This strategy is applied to screening of multiple clinical data sources for detection of individuals at risk of vital organ failure in partnership with a team at the University of British Columbia, St. Paul’s Hospital, NCE CECR Centre of Excellence for Prevention of Organ Failure (PROOF), and in collaboration with the James Hogg iCAPTURE Centre (Vancouver, BC). Individual patients matching failure risk patterns through the querying process are flagged as candidates for review and clinical decision making regarding treatment, medication, dosage, etc.
This use case describes an application of semantic technology for pre-symptomatic screening, detection, scoring and stratification of patients at risk of organ transplant failure due to immune rejection.
Experimental data from multiple OMICs modalities (genomics, proteomics) from a variety of sources (files, databases) are semantically integrated into a RDF-based network graph using user-supervised automated mapping and the dynamic building of an extensible applications ontology. During the import mapping process, thesauri are applied to harmonize synonyms and nomenclature differences between original data sources.
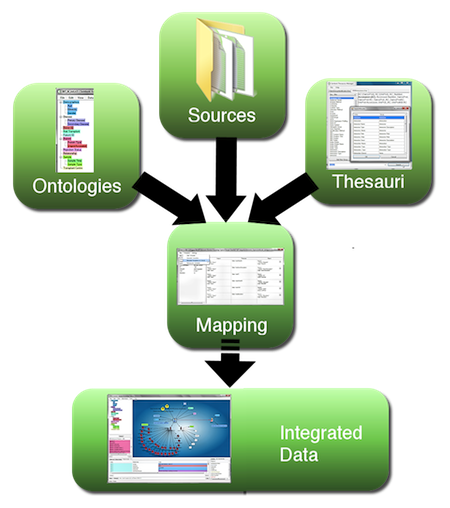
Figure 1. Workflow for semantic data integration, moving from heterogeneous data sources to integrated semantic data network. (A larger version of the figure is also available.)
Extraction, translation and loading (ETL) of data from files, instruments, images and diverse databases into a coherent environment still poses a major challenge, particularly if the data span across multiple fields of expertise and nomenclature. This is where semantic technology excels, as it builds on a common extensible data model (the Resource Description Framework, RDF), in which all data are represented in form of explicitly meaningful triples (A is related to B). As shown in Figure 1 above, data are mapped to a concept (this can be a dynamically built application ontology, a formal ontology from a public resource, or a combination of both) and diverse naming conventions are taken care of through application of one or multiple thesauri at the mapping phase, with further applications of inference within the semantic database. This assures uniform coherence while maintaining provenance and source accessibility. Integrating data coherently in context and being able to extend the data network dynamically is an example where applying semantic technologies makes the fundamentally required integration task much easier. Without these methods, integration would be far more difficult and time consuming to complete; and the result would be far less flexible.
Once data have been semantically integrated, several steps are involved to create a pattern, which can describe a biological process of interest and can be applied to new data for validation and screening. In this case, the pattern describes the likelihood of organ failure in transplant patients.
First, patterns that combine several biological indicators (potential molecular “biomarkers”) are identified by statistical analysis. To understand relevance and causality, these potential biomarkers are then mechanistically qualified by knowledge building methods. Insights gained from semantically joining findings from different clinical and experimental sources (medication data, co-morbidity data, transplantation data, gene expression data, proteomics data, etc. all maintained in different databases) allow researchers to better understand mechanistic aspects of biomarkers for organ failure at a functional level.
Next, the resulting data patterns of interest are captured and applied using semantic Visual SPARQL™ (SPARQL Protocol and RDF Query Language) technology. Domain-experienced users (not programmers or informaticians) can directly generate sophisticated queries by selecting nodes on the network graph and making elements of resulting pattern variable. This in turn will automatically generate a SPARQL query in the background without requiring any programmatic intervention or understanding of SPARQL from the user, so non-informatics researchers and clinicians can build complex queries, which reflect classifiers with functional biology characteristics. In this use case, SPARQL arrays combine genotypic and phenotypic protein expression information with range, weight, scoring and other filters to create and apply screening algorithms or “patterns”, sometimes also called “signatures”. SPARQL is uniquely well suited to sensitive and precise searches for data relationships across multiple, previously disconnected information sets, and is capable of detecting patterns within and between different data types and relationships even if the initial datasets are not formally joined under any common database schema. In practice, these query methods make it possible to apply complex screening algorithms across multiple data sources to deliver highly sensitive and specific patient screening, stratification and personalization outcomes.
Such patterns are placed in an Applied Semantic Knowledgebase (ASK™) for further validation and for decision support applications. ASK provides a collection of these patterns, applicable to screening and decision making. These may be applied to varied research foci and at the point of care, with data handling and protocols assisted by a complementary Process Manager software application.
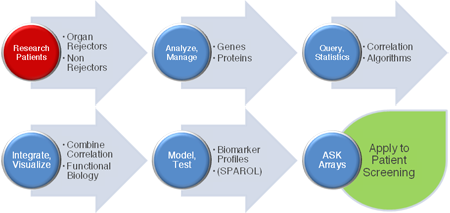
Figure 2. Workflow: moving from patient data to web-based combinatorial biomarker screening using ASK arrays (A larger version of the figure is also available.)
Using ASK makes it possible to actively screen previously disconnected, distributed datasets to identify and stratify results. This delivers applications suitable for sensitive, specific and informed decision making in Life Science research and clinical personalized medicine environments.
Other existing applications of ASK include hypothesis generation and testing, target profile creation and validation, compound efficacy and promiscuity screening, toxicity profiling and detection, treatment efficacy screening, and in this use case, screening of molecular patterns for risk of organ failure.
Numerous challenges must be overcome for decision support in personalized medicine. For example, patterns associated with common clinical endpoints such as disease states or treatment responses can include widely varied data, such as demographics, clinical symptoms and signs, imaging and laboratory results, therapeutics, and genetic, genomics, proteomics, and/or metabolomics data. Statistical correlations often find co-expressed data that are not necessarily or clearly functionally connected. Even if all data points result from the same disease state, co-varying data can represent very different biological processes and can exhibit the sum of multiple overlapping relationships.
Additionally, different data standards and definitions in laboratory and clinics have historically made merging of data into a common database or federated data model extremely difficult. To add to this challenge, the organizations that produce and/or maintain the data are autonomous entities. Independently collaborating institutions frequently change their data formats and database structures to fit their unique needs, most often for very good reasons. Data resources and descriptions frequently change in real-world settings.
Thus, more efficient and flexible methods for relationship consolidation, class hierarchy adjustment and resulting data integration are required to make inference, reasoning and rich screening for patterns of interest across research and clinical data sources practical and achievable. Semantic technologies are designed to address these challenges.
Gene transcription and protein expression levels associated with clinical endpoints (such as acute transplant rejection and non-rejection) were statistically identified, with robust correlation between independent analytical results. Results were merged into a semantic knowledge building framework to visualize and investigate associated correlations as well as mechanistic data relationships. To do this, significant elements were integrated within a correlation network and reduced to sub-networks of associated genes and proteins. Potential classifiers (biomarkers) from the correlation network were scaled using numerical properties (fold-change, p-value) to visually reduce network complexity and pre-select classifiers.
Next, functional biology references were added from published data sources. The biomarkers were further assessed according to mechanistic insights for biological relevance from the integrated network, to assure the combined classifiers made sense from a biological systems perspective. This step included import of parts of formal ontologies (in OWL and OBO) such as the Pathway Ontology from public resources (NCBO) as well as functional biology relevant references (NCBI's BioSystems, several Entrez databases, and SPARQL endpoint access to LODD) to qualify the validity of the model from a biological perspective with public knowledge. This makes it possible to understand the biomarker panels in context with their associated biological mechanisms. This step is important to researchers and regulatory agencies tasked with discovering and qualifying biomarker panels prior to their application in clinical settings.
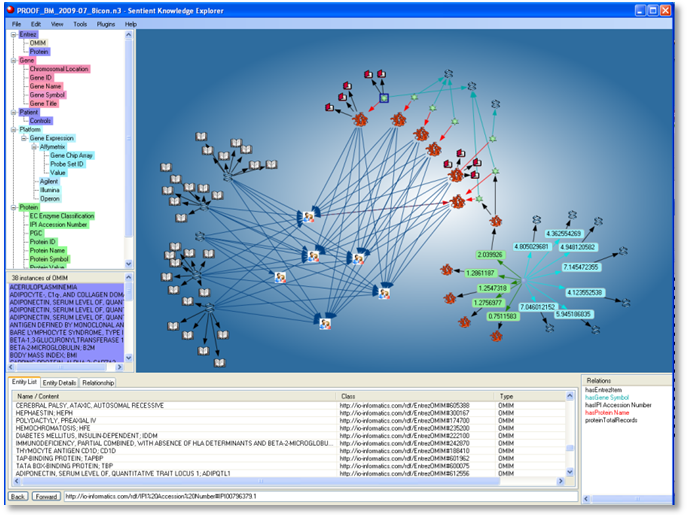
Figure 3. Resulting semantic data network: Public reference-enhanced experiments in a common ontology
Resulting sub-networks were identified as potential screening patterns (combined biomarker classifiers). Visual SPARQL queries were directly generated from interactive, user selected sub-networks without requiring knowledge of SPARQL. In Sentient, the Visual SPARQL capabilities have been extended to include filters on ranges, weighting, inclusion and exclusion criteria, etc. Sets of such SPARQL queries are captured and saved as arrays. These resulting arrays of screening models are applied and further validated with test cases and then applied to unknowns for screening. Models can be refined with newly confirmed cases to increase prediction precision.
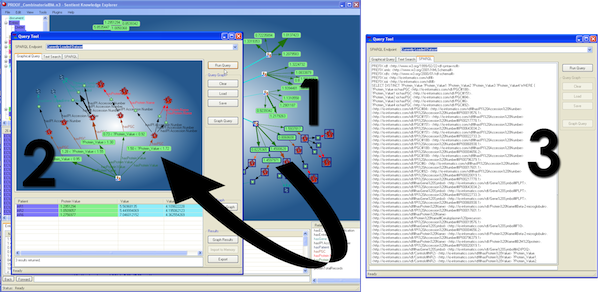
Figure 4. SPARQL creation directly from graph: Selecting nodes from the main network graph (1) generates a visual SPARQL representation of the query (2) and the actual SPARQL statement (3) automatically (a larger version of the figure is also available.)
For the clinician, simple web-based alerting of “hits” is provided, with scores to identify the closeness of fit of the patient to a risk pattern. Statistically supported scoring represents the confidence in the match (“hit-to-fit”) for informed decision-making.
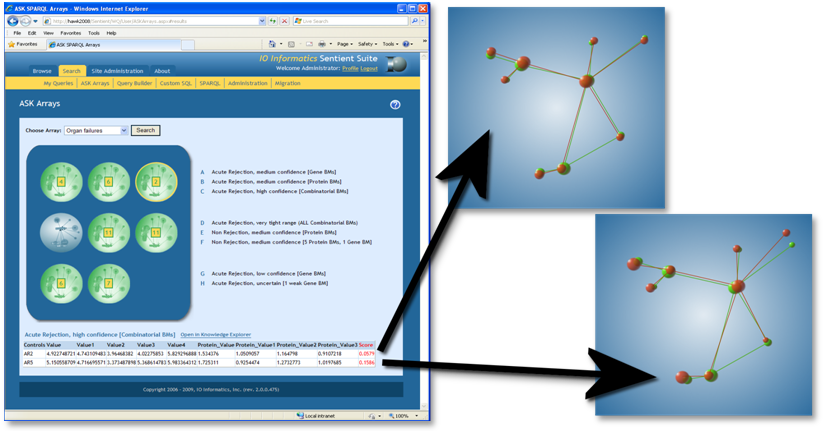
Figure 5. Web-browser accessible ASK arrays: Predictive screening as decision-support for organ failures (left: main interface with scoring) and “hit-to-fit” representation as decision support tool for goodness of prediction
An intuitive dashboard interface provides web-based access to clinicians, showing “hits” for patients at risk of organ failure or other important clinical events like immune rejection in the instance of transplants. Semantic technologies make it possible to run current, active searches across relevant data sources, to detect patterns indicating risk of organ failure. Complementary process management software is also in use at the PROOF Centre and iCAPTURE Centre sites and can be applied to send out cell phone alerts, text messages, etc., with associated dashboards.
Although value is widely recognized, combined molecular biomarker patterns are not yet commonly applied in production healthcare systems for personalized medicine, in part due to challenges outlined above. These challenges are addressed by technical advances outlined in the use case. In parallel, political, regulatory and commercialization challenges are rapidly turning into opportunities. PROOF Centre has engaged the FDA via voluntary data submission and is working towards FDA approval of these patterns for predictive, diagnostic, prescriptive and prognostic applications.
Tissue biopsies are currently the only way to properly monitor transplant patients for organ failure related to immune rejection. Biopsies are invasive but necessary to fine-tune the dosage of immunosuppressive drugs required by every transplant patient. Too small a dosage can result in organ rejection and potential organ failure; too much leaves patients susceptible to dangerous infections and eventual cancers. Biopsies are costly too; heart transplant patients undergo at least a dozen biopsies in the first year after transplant, at a composite cost of $5,000- $10,000 each.1
The ability to predict risk of organ failure with a simple, inexpensive blood test and screen using ASK significantly reduces the need for biopsies, while improving timeliness, sensitivity and specificity of screens for organ failure risk2. This approach reduces time and cost burdens on the healthcare system and improves safety and quality of life for patients. A similar statement can be made with regards to improved diagnosis of the actual occurrence of rejection through diagnostic biomarker panels, whereupon the biopsy may ultimately be replaceable.
In Healthcare and Life Sciences domains, broadly integrated semantic knowledge applications are well suited to closing the loop at multiple points in the life cycle of research: to improve outcomes for drug discovery, development, screening and personalized treatment.
© Copyright 2011, IO Informatics, Inc., and University British Columbia (UBC),