HCLSIG/SWANSIOC/Actions/RhetoricalStructure/alignment/mediumgrain
Goal: Discuss and align medium grain ontologies for biomedical paper structure
DRO = "Document Rhetoric Ontology"
Procedure (modified from Tim's suggestion):
- ORB + DRO first. (ORB + DRO = DRO* ). I think this is important to do first because it is so simple. ORB has also had the most discussion, very well-grounded in use cases. And there is another reason to do ORB + DRO first...
- DRO* + Data-Experiment (DEXI) next. (DRO* + DEXI = DRO**). Why? Because
- DEXI has already been under construction for a year, and includes a lot of grounding and cross group discussion, so it is quite well-grounded; and
- DRO and DEXI have a definite and strong overlap, that can impact on details further on, when you bring along DOCO.
- Align with Body section of Anita's model
- DRO** + DOCO.
In general: Align the most firmly-grounded (in both use cases and inter-group discussions) and simplest ontologies first, because they are more foundational and straightforward.
Upcoming discussions
Gully Burns, Sudesna Das, David Newman, Jodi Schneider - discussion about medium-grained and KEfED/DExI/myExperiment
Relevant ontologies
Coarse-grained Ontology of Rhetorical Blocks
To align
- DRO, starting with Middle-grained document structure
- DEXI: current OWL (old image) (HCLSIG/SWANSIOC/Actions/SWANmyExpArray#Ontology older info)
- DoCO, see listserv message from David Shotton 2010-11-12 and 2010-11-15 discussion
Use case
- Medium Grain use case is Improving the Structure of Digital Publications in the Computer Science Domain
Past discussions
2011-04-14 HCLS meeting
Howard: It seems like either paragraph or figure could serve a variety of semantic or rhetorical roles.
Tim: Blocks and [clauses?]??? play different rhetorical roles in the document, so you may want to annotate them differently.
Anita: I thought that "fine-grained" was what SWAN annotated: recitation, evidence, claims.
You can't always point to a single place where a claim is stated. It may be stated several places, and narrative-focused.
So is the challenge: how to connect fine-grained *process* markup to fine-grained *document* markup?
Gully: There's a difference between the scientific process and the construction of argument across documents.
In Slide 11 [on SWAN] -- a lot of elements could be distinguished into categories It could be useful to categorize the elements of as
- Intra-document
- Intra-experiment
- Extra-document
Even within a document, there may be cross-experiment argumentation.
Science draws from an external set of knowledge -- to provide motivation, interpretation, experimental design.
There are also intra-document relationships -- like pointing to figures, or constructing an experimental design that is specifically geared towards a certain question.
Further, a document may contain multiple experiments.
Tim: Science is "warranted true belief". There are two kinds of warrant: either a reference to someone else's papers (external evidence) or reference to the experiment - details, reasoning, data etc. (internal evidence relying on the provenance and logical procedures of the papers).
Gully: Tim - I think my point was just that the slides have these different types of relations in one place. I'd like to categorize these types, separating extra-document knowledge from intra-document evidence and process.
At ISI, the machine-reading group works on this problem, they distinguish between the "reading machine" and the "reasoning machine": what do we annotate the text with; how do we use those annotations to reason with?
There are process elements and semantic representation. For discourse (on the process side) -- what are the general aspects of what we're trying to annotate, in order to serve the purpose of reasoning? We want to find the underlying reasoning and do computation over it.
Anita: The question "what are we modelling?" needs to be proceeded by "why are we modelling?". Do we need new use cases?
The existing use cases are to:
Create a claim-evidence network spanning over documents}
- Provide templates for authoring of documents (Use Case #2: Publication authoring here)
- Link publications and enhance search
Jodi: Integration between the paper and the science is the main point here - so how do you get claims out of a single paper? Anita: and link them to the underlying experimental model?
Gully: The use cases we are talking about are informatics use cases, but we should be focusing on biologists use cases. Can we suggest to an biologist end-user what a new field or direction would be? To make a claim network useful in practice, by taking them to a place where they haven't worked before?
Tim: great point Gully - also point them to experiments and tools to do that work?
Alex: when biologists go to libraries, they have a particular problem, are following a protocol, want to find papers where same protocol is used with slight modifications - not so much claims.
Gully: The ultimate goal is for scientist to be able to design a new experiment based on the knowledge you have based your model on. We want machines that make a prediction for the scientist, and propose a new experiment. So pull out all claims about observations and interpretations, munge them together into a model that you can do reasoning on, and generate an experimental design to test the new hypothesis.
Gully: Hanalyzer, from UC Denver, is an interesting system to consider. Related video
They work in a well-defined domain which they understand well, and besides the tools, they have a human analyst to "think about the science". This is a good recipe for success: solve problems for a specific domain.
There are two main differences from what we are doing: first, it is a complete system (with user interfaces) NOT just an ontology. Second, it is specific, whereas what we want is very general
Joanne: So how do you bridge perspectives, between a domain and external [general?] perspectives? You need to know what role you are in. Different types of evidence are used in different fields.
Next steps from Anita: 1) investigate workshop 2) write new use case 3) Joanne to invite someone from the Hunter group
Other possible next-steps: presentation from the ISI machine-reading group.
Other thoughts to take up: Howard: Not sure stories or narratives always strictly follow beginning-middle-end series: roles often shift about with lots of non-linear reflection and impact out-of-sequence.
2011-01-31 HCLS meeting
2010-12-14 at 9 AM EST
Sudeshna Das & Jodi Schneider about DExI
Sudeshna shared File:DeXI-intro.pdf, which include the latest DExI draft (below). The overall goal is to tie discourse in with the experiment (which consists of data, metadata, and computation). Discourse is in the results section (claims made) and in the discussion section. Link claims to data + experiment. "Lego blocks" of the various elements.
DExI draft v5 2010-11-01 (png): 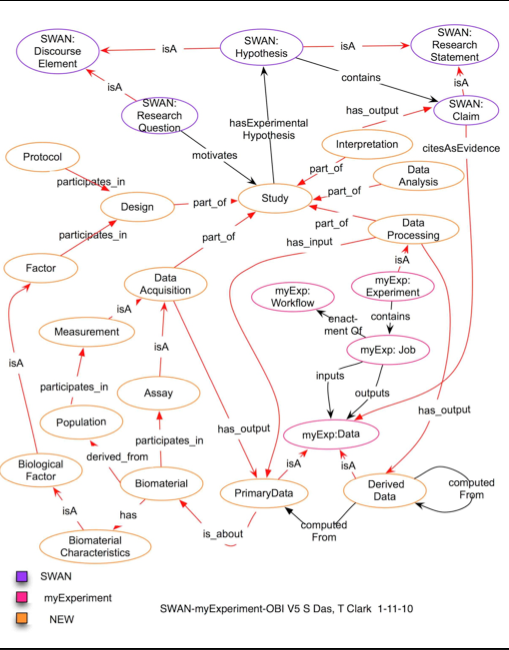
Design
Design includes cases, controls, protocols, hypothesis, what is going to be measured. Depends on the type of experiment: FDA requires thorough design for clinical/human studies. Could also be animal studies, cell lines. Design may be observational or intervention-based (doing something to one cohort, to observe the differences). The variables are the factors of interest.
(Data) Acquisition
Primary data
input: biomaterials, population output: raw data
Data Processing
data is processed/transformed
Data Analysis
OBI doesn't have one single class for this.
Interpretation
"these steps (that) make sense to biomedical researchers" 1 outcome variable influencing variables (that influence their outcome) an information artifact - data collection, measurement, recording, assays
Various approaches to interpretation, for instance, Gully distinguishes between observational and interpretive claims.
Use Cases for DExI
- interoperability of experiment repositories, Science Collaboration Framework, Stephane C. -- in Drupal
- ISA-tab -- Susannah's group at Oxford -- bii (generalised biomedical data)
- link claims and publication to experiments, using AO
- interoperability between repositories (e.g. ISA-tab, PDexpression) -- want to query against both repositories. Name-value pair (semantics are too complicated) -- see slide 12
2010-12-07 at 10 AM EST/3 PM GMT
Tim Clark, Silvio Peroni, Jodi Schneider, David Shotton, Anita de Waard. (Alexander Garcia Castro -- unfortunately with line trouble)
Anita gave a little history about the middle-grained ontology project, which we're calling DRO.
We then discussed ORB, which consists of 5 disjoint classes: Header Introduction Methods Discussion References Some discussion concerned its alignment with external ontologies and structure, which we passed on to Tim to convey (and has since been discussed on the email list).
We then discussed DeXI, which is subset of OBI+MyExperiment+SWAN, under development by Sudeshna Das and others. We've since learned that the current version is at http://swan-ontology.googlecode.com/svn/trunk/experiments.owl
Tim explained that they're starting with genomic lab experiments and observational population studies. Covers experimental hypotheses, design, factors (similar to DRO's objects of study), characteristics of the biomaterial. While it's still missing a few concepts from the computational side, it already gives the ability to take an experimental finding in a paper and map it to the things you could export from arrayExpress (a public repository for microarray gene expression data, "genBank for arrays").
We next discussed DoCO. David Shotton explained that its goal is to republish documents and metadata as open Linked Data, starting with documents in the NLM DTD, to support the Open Bibliographic Project. There was some skepticism regarding converting a DTD into RDF. Smaller publishers are interested in using this; larger publishers have little incentive to change DTDs and have established workflows around custom formats used for a great variety of structures (not just research articles but also books, etc).
Outcomes:
- Tim will ping Paolo about the ORB write-up for a W3C note
- Anita will share Ron Daniel's email about aligning the ORB Header with OAI, PRISM, DC, ...
- Tim to put Jodi in touch with Sudeshna Das regarding DeXI
- Tim to take feedback about ORB to Paolo and Tudor (e.g. referencing Head and References externally, Tail, …)
2010-11-23
Anita, Alex Garcia and Jodi discussed the possibilities for alignment between DoCO and Medium-grained document structure. DoCO is currently being developed as part of SPAR (SPAR intro).
Our general conclusion was that David Shotton's proposal [particularly, it's here: http://lists.w3.org/Archives/Public/public-semweb-lifesci/2010Nov/att-0040/Shotton_discussion_paper_on_DRO.pdf in this PDF] (attached to David Shotton's listserv message 2010-11-17 was on target. However, we want to:
- use existing ontologies for references (BIBO? ...?)
- use existing ontologies for the header (PRISM? DC?...?)
- check the use of fabio: (e.g. for Experimental Protocol)
- check the use dro:
- check the use of sro:
A few questions regarding DoCO came up. The combination of document components, rhetorical components, rhetorical blocks, and structural patterns confused us. We expected these to be several smaller ontologies. Another question (David, perhaps you can answer this) is why you prefer imports into a larger ontology, as opposed to building an application profile? Rhetorical components, for instance, may already be handled adequately by SALT, SWAN, and ScholOnto.
We also discussed whether we wanted to get beyond ontologies to also address authoring and/or textmining (with a schema or DTDs drawing from ontologies). Anita pointed out that in our 3 use cases, 1 involves authoring. Further, for authoring we're limited to continguous sections (as opposed to post-hoc rhetorical component detection).